WEBNETListen to the readings for May 29 from the NET Bible (Video)
or the World English Bible
This page is now calculating which verses have comments and referenced comments. Please be patient.
You will not be able to view any Comments in the Verse sections below until this notice disappears
Notes that are not verse specific
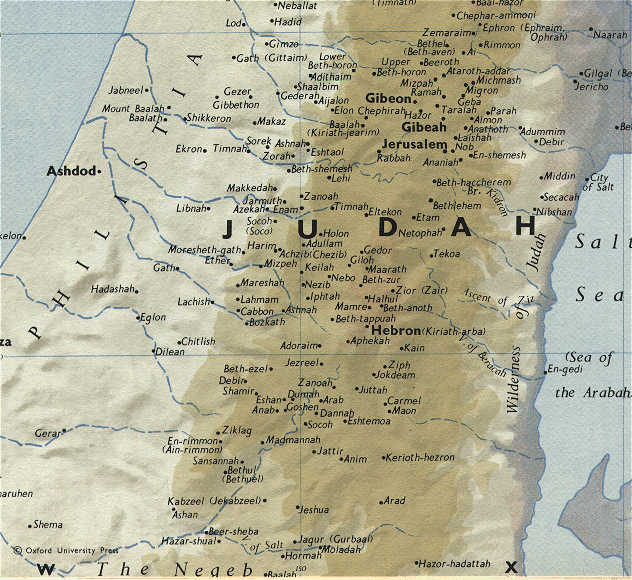
You will need a map for this. Try this one!
Peter [UK] Comment added in 2001 Reply to Peter
When considering Judah, it should be remembered that the tribal allotment of Simeon lay within its (Judah's) boundaries (Josh 19:1).
Michael Parry [Montreal (Can)] Comment added in 2004 Reply to Michael
To view any notes specific to the verses below, use the icons:
 Comments made directly on this verse
Comments made directly on this verse  Comments made elsewhere with reference to this verse
Comments made elsewhere with reference to this verse
Please note that comments referring to a range of verses will be generated for every verse in that range
1. This then was the lot <1486> of the tribe <4294> of the children <1121> of Judah <3063> by their families <4940>; even to the border <1366> of Edom <123> the wilderness <4057> of Zin <6790> southward <5045> was the uttermost part <7097> of the south coast <8486>.
2. And their south <5045> border <1366> was from the shore <7097> of the salt <4417> sea <3220>, from the bay <3956> that looketh <6437>(8802) southward <5045>:
3. And it went out <3318>(8804) to the south side <5045> to Maalehacrabbim <4610>, and passed <5674>(8804) along to Zin <6790>, and ascended up <5927>(8804) on the south side <5045> unto Kadeshbarnea <6947>, and passed <5674>(8804) along to Hezron <2696>, and went up <5927>(8804) to Adar <146>, and fetched a compass <5437>(8738) to Karkaa <7173>:
4. From thence it passed <5674>(8804) toward Azmon <6111>, and went out <3318>(8804) unto the river <5158> of Egypt <4714>; and the goings out <8444> of that coast <1366> were at the sea <3220>: this shall be your south <5045> coast <1366>.
5. And the east <6924> border <1366> was the salt <4417> sea <3220>, even unto the end <7097> of Jordan <3383>. And their border <1366> in the north <6828> quarter <6285> was from the bay <3956> of the sea <3220> at the uttermost part <7097> of Jordan <3383>:
6. And the border <1366> went up <5927>(8804) to Bethhogla <1031>, and passed <5674>(8804) along by the north <6828> of Betharabah <1026>; and the border <1366> went up <5927>(8804) to the stone <68> of Bohan <932> the son <1121> of Reuben <7205>:
7. And the border <1366> went up <5927>(8804) toward Debir <1688> from the valley <6010> of Achor <5911>, and so northward <6828>, looking <6437>(8802) toward Gilgal <1537>, that is before <5227> the going up <4608> to Adummim <131>, which is on the south side <5045> of the river <5158>: and the border <1366> passed <5674>(8804) toward the waters <4325> of Enshemesh <5885>, and the goings out <8444> thereof were at Enrogel <5883>:
8. And the border <1366> went up <5927>(8804) by the valley <1516> of the son <1121> of Hinnom <2011> unto the south <5045> side <3802> of the Jebusite <2983>; the same is Jerusalem <3389>: and the border <1366> went up <5927>(8804) to the top <7218> of the mountain <2022> that lieth before <6440> the valley <1516> of Hinnom <2011> westward <3220>, which is at the end <7097> of the valley <6010> of the giants <7497> northward <6828>:
9. And the border <1366> was drawn <8388>(8804) from the top <7218> of the hill <2022> unto the fountain <4599> of the water <4325> of Nephtoah <5318>, and went out <3318>(8804) to the cities <5892> of mount <2022> Ephron <6085>; and the border <1366> was drawn <8388>(8804) to Baalah <1173>, which is Kirjathjearim <7157>:
10. And the border <1366> compassed <5437>(8738) from Baalah <1173> westward <3220> unto mount <2022> Seir <8165>, and passed <5674>(8804) along unto the side <3802> of mount <2022> Jearim <3297>, which is Chesalon <3693>, on the north side <6828>, and went down <3381>(8804) to Bethshemesh <1053>, and passed on <5674>(8804) to Timnah <8553>:
11. And the border <1366> went out <3318>(8804) unto the side <3802> of Ekron <6138> northward <6828>: and the border <1366> was drawn <8388>(8804) to Shicron <7942>, and passed along <5674>(8804) to mount <2022> Baalah <1173>, and went out <3318>(8804) unto Jabneel <2995>; and the goings out <8444> of the border <1366> were at the sea <3220>.
12. And the west <3220> border <1366> was to the great <1419> sea <3220>, and the coast <1366> thereof. This is the coast <1366> of the children <1121> of Judah <3063> round about <5439> according to their families <4940>.
13. And unto Caleb <3612> the son <1121> of Jephunneh <3312> he gave <5414>(8804) a part <2506> among <8432> the children <1121> of Judah <3063>, according <413> to the commandment <6310> of the LORD <3068> to Joshua <3091>, even the city <7151> of Arba <704>(8677) <7153> the father <1> of Anak <6061>, which city is Hebron <2275>.
14. And Caleb <3612> drove <3423>(8686) thence the three <7969> sons <1121> of Anak <6061>, Sheshai <8344>, and Ahiman <289>, and Talmai <8526>, the children <3211> of Anak <6061>.
15. And he went up <5927>(8799) thence to the inhabitants <3427>(8802) of Debir <1688>: and the name <8034> of Debir <1688> before <6440> was Kirjathsepher <7158>.
16. And Caleb <3612> said <559>(8799), He that smiteth <5221>(8686) Kirjathsepher <7158>, and taketh <3920>(8804) it, to him will I give <5414>(8804) Achsah <5915> my daughter <1323> to wife <802>.
17. And Othniel <6274> the son <1121> of Kenaz <7073>, the brother <251> of Caleb <3612>, took <3920>(8799) it: and he gave <5414>(8799) him Achsah <5915> his daughter <1323> to wife <802>.
18. And it came to pass, as she came <935>(8800) unto him, that she moved <5496>(8686) him to ask <7592>(8800) of her father <1> a field <7704>: and she lighted off <6795>(8799) her ass <2543>; and Caleb <3612> said <559>(8799) unto her, What wouldest thou?
19. Who answered <559>(8799), Give <5414>(8798) me a blessing <1293>; for thou hast given <5414>(8804) me a south <5045> land <776>; give <5414>(8804) me also springs <1543> of water <4325>. And he gave <5414>(8799) her the upper <5942> springs <1543>, and the nether <8482> springs <1543>.
20. This is the inheritance <5159> of the tribe <4294> of the children <1121> of Judah <3063> according to their families <4940>.
21. And the uttermost <7097> cities <5892> of the tribe <4294> of the children <1121> of Judah <3063> toward the coast <1366> of Edom <123> southward <5045> were Kabzeel <6909>, and Eder <5740>, and Jagur <3017>,
22. And Kinah <7016>, and Dimonah <1776>, and Adadah <5735>,
23. And Kedesh <6943>, and Hazor <2674>, and Ithnan <3497>,
24. Ziph <2128>, and Telem <2928>, and Bealoth <1175>,
25. And Hazor <2674>, Hadattah <2675>, and Kerioth <7152>, and Hezron <2696>, which is Hazor <2674>,
26. Amam <538>, and Shema <8090>, and Moladah <4137>,
27. And Hazargaddah <2693>, and Heshmon <2829>, and Bethpalet <1046>,
28. And Hazarshual <2705>, and Beersheba <884>, and Bizjothjah <964>,
29. Baalah <1173>, and Iim <5864>, and Azem <6107>,
30. And Eltolad <513>, and Chesil <3686>, and Hormah <2767>,
31. And Ziklag <6860>, and Madmannah <4089>, and Sansannah <5578>,
32. And Lebaoth <3822>, and Shilhim <7978>, and Ain <5871>, and Rimmon <7417>: all the cities <5892> are twenty <6242> and nine <8672>, with their villages <2691>:
33. And in the valley <8219>, Eshtaol <847>, and Zoreah <6881>, and Ashnah <823>,
34. And Zanoah <2182>, and Engannim <5873>, Tappuah <8599>, and Enam <5879>,
35. Jarmuth <3412>, and Adullam <5725>, Socoh <7755>, and Azekah <5825>,
36. And Sharaim <8189>, and Adithaim <5723>, and Gederah <1449>, and Gederothaim <1453>; fourteen <702> <6240> cities <5892> with their villages <2691>:
37. Zenan <6799>, and Hadashah <2322>, and Migdalgad <4028>,
38. And Dilean <1810>, and Mizpeh <4708>, and Joktheel <3371>,
39. Lachish <3923>, and Bozkath <1218>, and Eglon <5700>,
40. And Cabbon <3522>, and Lahmam <3903>, and Kithlish <3798>,
41. And Gederoth <1450>, Bethdagon <1016>, and Naamah <5279>, and Makkedah <4719>; sixteen <8337> <6240> cities <5892> with their villages <2691>:
42. Libnah <3841>, and Ether <6281>, and Ashan <6228>,
43. And Jiphtah <3316>, and Ashnah <823>, and Nezib <5334>,
44. And Keilah <7084>, and Achzib <392>, and Mareshah <4762>; nine <8672> cities <5892> with their villages <2691>:
45. Ekron <6138>, with her towns <1323> and her villages <2691>:
46. From Ekron <6138> even unto the sea <3220>, all that lay near <3027> Ashdod <795>, with their villages <2691>:
47. Ashdod <795> with her towns <1323> and her villages <2691>, Gaza <5804> with her towns <1323> and her villages <2691>, unto the river <5158> of Egypt <4714>, and the great <1419>(8675) <1366> sea <3220>, and the border <1366> thereof:
48. And in the mountains <2022>, Shamir <8069>, and Jattir <3492>, and Socoh <7755>,
49. And Dannah <1837>, and Kirjathsannah <7158>, which is Debir <1688>,
50. And Anab <6024>, and Eshtemoh <851>, and Anim <6044>,
51. And Goshen <1657>, and Holon <2473>, and Giloh <1542>; eleven <259> <6240> cities <5892> with their villages <2691>:
52. Arab <694>, and Dumah <1746>, and Eshean <824>,
53. And Janum <3241>, and Bethtappuah <1054>, and Aphekah <664>,
54. And Humtah <2547>, and Kirjatharba <7153>, which is Hebron <2275>, and Zior <6730>; nine <8672> cities <5892> with their villages <2691>:
55. Maon <4584>, Carmel <3760>, and Ziph <2128>, and Juttah <3194>,
56. And Jezreel <3157>, and Jokdeam <3347>, and Zanoah <2182>,
57. Cain <7014>, Gibeah <1390>, and Timnah <8553>; ten <6235> cities <5892> with their villages <2691>:
58. Halhul <2478>, Bethzur <1049>, and Gedor <1446>,
59. And Maarath <4638>, and Bethanoth <1042>, and Eltekon <515>; six <8337> cities <5892> with their villages <2691>:
60. Kirjathbaal <7154>, which is Kirjathjearim <7157>, and Rabbah <7237>; two <8147> cities <5892> with their villages <2691>:
61. In the wilderness <4057>, Betharabah <1026>, Middin <4081>, and Secacah <5527>,
62. And Nibshan <5044>, and the city of Salt <5898>, and Engedi <5872>; six <8337> cities <5892> with their villages <2691>.
63. As for the Jebusites <2983> the inhabitants <3427>(8802) of Jerusalem <3389>, the children <1121> of Judah <3063> could <3201>(8804)(8675) <3201>(8799) not drive them out <3423>(8687): but the Jebusites <2983> dwell <3427>(8799) with the children <1121> of Judah <3063> at Jerusalem <3389> unto this day <3117>.
Notes that are not verse specific
ch.20 - WHAT DO YOU RELY ON?
Sometimes the things we think are the most stable things in our lives are suddenly taken away from us and we find our lives falling to pieces around us. We can often find that we place too much emphasis on possessions, money, jobs, family, relationships or enjoying life that we end up making one or more of them the foundation of our life. If those sort of things become our foundation our whole world can fall down when they fail. There were people in Isaiah 20 who trusted in Cush and Egypt and in their military strength to deliver them from the might of Assyria. But the strength of Egypt and Cush were to fail and those who boasted in their strength were to be afraid and put to shame. There is only one thing that is really strong enough and secure enough for us to place our faith and strength in. That strength is in the LORD our God. He will never change. He will always be strong. He will always be there for us and he will never let us down. So even if the world moves and shifts around us as shifting sands, we can rely on the rock that we have built on - God our stronghold.
Robert Prins [Auckland - Pakuranga - (NZ)] Comment added in 2002 Reply to Robert
Isa 20.
What, if any, is the significance to the 3 years time period Isaiah walked around stripped and barefoot?
The following ideas are mostly from bro. Harry Whittaker's book Isaiah. On the face of it, this is a prophecy with a specific short-term fulfillment, sandwiched between 2 blocks of 7 "burdens" (Isa. ch. 15-23), all of which refer to nations surrounding Judah. It would appear then that this short-term prophecy, readily put to the test within 3 years, served to guarantee the accuracy of the other burdens, both in their primary and later fulfillments.
Ashdod, formerly a Philistine city, had likely become a Jewish fortress. And at this time there was a lean-on-Egypt policy (Isaiah would later denounce this!) to the point where Egyptian help took the form of a garrison which essentially made Ashdod an Egyptian outpost. But now, through what Isaiah was commanded by God to do, those Egyptians would see the shame of captivity that would soon happen to them.
At this time Egypt was dominated by an Ethiopian dynasty. So if in this soon coming encounter, mighty Egypt and Ethiopia are to be proved so futile and worthless a pair of allies, due to Assyrian's invasion and conquest of them, what hope is there for Judah when Assyrian expansion comes to full blood? "How shall we escape?" That is, if pagan nations with so little light suffer such retributions from God, what can His faithless chosen people expect form Assyria, the rod of God's anger (Isa 10:5)?
Is it possible then that in days soon to come, modern Israel, trying to be friends with Egypt, will similarly pay for its faithless misguided statesmanship and be brought to a humiliation (but also deliverance!) comparable to that which came about in Isaiah's day?
Wes Booker [South Austin Texas USA] Comment added in 2013 Reply to Wes
To view any notes specific to the verses below, use the icons:
 Comments made directly on this verse
Comments made directly on this verse  Comments made elsewhere with reference to this verse
Comments made elsewhere with reference to this verse
Please note that comments referring to a range of verses will be generated for every verse in that range
1. In the year <8141> that Tartan <8661> came <935>(8800) unto Ashdod <795>, (when Sargon <5623> the king <4428> of Assyria <804> sent <7971>(8800) him,) and fought <3898>(8735) against Ashdod <795>, and took <3920>(8799) it;
2. At the same time <6256> spake <1696>(8765) the LORD <3068> by <3027> Isaiah <3470> the son <1121> of Amoz <531>, saying <559>(8800), Go <3212>(8798) and loose <6605>(8765) the sackcloth <8242> from off thy loins <4975>, and put off <2502>(8799) thy shoe <5275> from thy foot <7272>. And he did so <6213>(8799), walking <1980>(8800) naked <6174> and barefoot <3182>.
3. And the LORD <3068> said <559>(8799), Like as my servant <5650> Isaiah <3470> hath walked <1980>(8804) naked <6174> and barefoot <3182> three <7969> years <8141> for a sign <226> and wonder <4159> upon Egypt <4714> and upon Ethiopia <3568>;
4. So shall the king <4428> of Assyria <804> lead away <5090>(8799) the Egyptians <4714> prisoners <7628>, and the Ethiopians <3568> captives <1546>, young <5288> and old <2205>, naked <6174> and barefoot <3182>, even with their buttocks <8357> uncovered <2834>(8803), to the shame <6172> of Egypt <4714>.
5. And they shall be afraid <2865>(8804) and ashamed <954>(8804) of Ethiopia <3568> their expectation <4007>, and of Egypt <4714> their glory <8597>.
6. And the inhabitant <3427>(8802) of this isle <339> shall say <559>(8804) in that day <3117>, Behold, such <3541> is our expectation <4007>, whither we flee <5127>(8804) for help <5833> to be delivered <5337>(8736) from <6440> the king <4428> of Assyria <804>: and how shall we escape <4422>(8735)?
1. The burden <4853> of the desert <4057> of the sea <3220>. As whirlwinds <5492> in the south <5045> pass <2498>(8800) through; so it cometh <935>(8802) from the desert <4057>, from a terrible <3372>(8737) land <776>.
2. A grievous <7186> vision <2380> is declared <5046>(8717) unto me; the treacherous dealer <898>(8802) dealeth treacherously <898>(8802), and the spoiler <7703>(8802) spoileth <7703>(8802). Go up <5927>(8798), O Elam <5867>: besiege <6696>(8798), O Media <4074>; all the sighing <585> thereof have I made to cease <7673>(8689).
3. Therefore are my loins <4975> filled <4390>(8804) with pain <2479>: pangs <6735> have taken hold <270>(8804) upon me, as the pangs <6735> of a woman that travaileth <3205>(8802): I was bowed down <5753>(8738) at the hearing <8085>(8800) of it; I was dismayed <926>(8738) at the seeing <7200>(8800) of it.
4. My heart <3824> panted <8582>(8804), fearfulness <6427> affrighted <1204>(8765) me: the night <5399> of my pleasure <2837> hath he turned <7760>(8804) into fear <2731> unto me.
5. Prepare <6186>(8800) the table <7979>, watch <6822>(8800) in the watchtower <6844>, eat <398>(8800), drink <8354>(8800): arise <6965>(8798), ye princes <8269>, and anoint <4886>(8798) the shield <4043>.
6. For thus hath the Lord <136> said <559>(8804) unto me, Go <3212>(8798), set <5975>(8685) a watchman <6822>(8764), let him declare <5046>(8686) what he seeth <7200>(8799).
7. And he saw <7200>(8804) a chariot <7393> with a couple <6776> of horsemen <6571>, a chariot <7393> of asses <2543>, and a chariot <7393> of camels <1581>; and he hearkened <7181>(8689) diligently <7182> with much <7227> heed <7182>:
8. And he cried <7121>(8799), A lion <738>: My lord <136>, I stand <5975>(8802) continually <8548> upon the watchtower <4707> in the daytime <3119>, and I am set <5324>(8737) in my ward <4931> whole nights <3915>:
9. And, behold, here cometh <935>(8802) a chariot <7393> of men <376>, with a couple <6776> of horsemen <6571>. And he answered <6030>(8799) and said <559>(8799), Babylon <894> is fallen <5307>(8804), is fallen <5307>(8804); and all the graven images <6456> of her gods <430> he hath broken <7665>(8765) unto the ground <776>.
10. O my threshing <4098>, and the corn <1121> of my floor <1637>: that which I have heard <8085>(8804) of the LORD <3068> of hosts <6635>, the God <430> of Israel <3478>, have I declared <5046>(8689) unto you.
11. The burden <4853> of Dumah <1746>. He calleth <7121>(8802) to me out of Seir <8165>, Watchman <8104>(8802), what of the night <3915>? Watchman <8104>(8802), what of the night <3915>?
12. The watchman <8104>(8802) said <559>(8804), The morning <1242> cometh <857>(8804), and also the night <3915>: if ye will enquire <1158>(8799), enquire <1158>(8798) ye: return <7725>(8798), come <857>(8798).
13. The burden <4853> upon Arabia <6152>. In the forest <3293> in Arabia <6152> shall ye lodge <3885>(8799), O ye travelling companies <736> of Dedanim <1720>.
14. The inhabitants <3427>(8802) of the land <776> of Tema <8485> brought <857>(8689) water <4325> to him <7125>(8800) that was thirsty <6771>, they prevented <6923>(8765) with their bread <3899> him that fled <5074>(8802).
15. For they fled <5074>(8804) from <6440> the swords <2719>, from <6440> the drawn <5203>(8803) sword <2719>, and from <6440> the bent <1869>(8803) bow <7198>, and from <6440> the grievousness <3514> of war <4421>.
16. For thus hath the Lord <136> said <559>(8804) unto me, Within a year <8141>, according to the years <8141> of an hireling <7916>, and all the glory <3519> of Kedar <6938> shall fail <3615>(8804):
17. And the residue <7605> of the number <4557> of archers <7198>, the mighty men <1368> of the children <1121> of Kedar <6938>, shall be diminished <4591>(8799): for the LORD <3068> God <430> of Israel <3478> hath spoken <1696>(8765) it.
Notes that are not verse specific
TEACHING
One thing I have learned through teaching the word of God to various people is that the person who has benefited most and come to the fullest understanding from what I have taught, has been myself. It never ceases to amaze me how that teaching something to someone else reinforces what you already know. So Paul's advice to Philemon is absolutely brilliant! He says, "I pray that you may be active in sharing your faith, so that you will have a full understanding of every good thing we have in Christ."
So we learn that there are, not two, but three really good reasons to share our faith. We need to share our faith to spread the gospel message so that others can be saved. We need to do it to obey the commandment that Christ gave. And we should share our faith so that we can consolidate and strengthen our own faith as we obey Christ's commandment and spread the gospel.
Of course, Paul's advice was not just for Philemon, it is for us as well. So let us not just share our faith, but be active in sharing our faith. Let's do it more and more. Paul prayed it for you too, "I pray that you may be active in sharing your faith, so that you will have a full understanding of every good thing we have in Christ."
Robert Prins [Auckland - Pakuranga - (NZ)] Comment added in 2002 Reply to Robert
Philemon had been the master of the runaway slave Onesimus. (Yes, slaves were kept even by Brothers and Sisters in Christ.) Just spare a moment today to try and work out the events which led to Onesimus agreeing to go back to Colosse, and actually to Brother Philemon’s house. Perhaps it was something like this: Paul, in prison, was brought a runaway slave who wanted to know about Jesus. He taught him the Truth, and Onesimus was baptised. Paul then wrote the letter to Colosse, and asked Onesimus a huge favour – “Will YOU kindly deliver this for me?”
“To Colosse? To Philemon? Well, I …er …” Whatever discussion took place Onesimus agreed, probably because Paul had given him a letter addressed to Brother Philemon. “Give him this,” Paul encouraged, “and I’m sure you'll be all right.” And, we have to suppose, he was.
David Simpson [Worcester (UK)] Comment added in 2007 Reply to David
Philemon (means friendly) was the master and Onesimus (means profitable) was the servant (slave). Under Roman law a runaway slave could be sentenced to death. However, Philemon took Onesimus back, not only as a servant but also as a brother. The parallel exists between our master, Jesus, and us as servants. If we remain faithful we are called friends (John 15:14). If we stray and come back we are welcomed (Luke 15:32). If we stay away, we are subject to death as the Roman slave would be. Nevertheless, we should realize that our service, unlike Onesimus', is considered unprofitable (Luke 17:10).
Michael Parry [Montreal (Can)] Comment added in 2009 Reply to Michael
To view any notes specific to the verses below, use the icons:
 Comments made directly on this verse
Comments made directly on this verse  Comments made elsewhere with reference to this verse
Comments made elsewhere with reference to this verse
Please note that comments referring to a range of verses will be generated for every verse in that range
1. Paul <3972>, a prisoner <1198> of Jesus <2424> Christ <5547>, and <2532> Timothy <5095> our brother <80>, unto Philemon <5371> our <2257> dearly beloved <27>, and <2532> fellowlabourer <4904>,
2. And <2532> to our beloved <27> Apphia <682>, and <2532> Archippus <751> our <2257> fellowsoldier <4961>, and <2532> to the church <1577> in <2596> thy <4675> house <3624>:
3. Grace <5485> to you <5213>, and <2532> peace <1515>, from <575> God <2316> our <2257> Father <3962> and <2532> the Lord <2962> Jesus <2424> Christ <5547>.
4. I thank <2168>(5719) my <3450> God <2316>, making <4160>(5734) mention <3417> of thee <4675> always <3842> in <1909> my <3450> prayers <4335>,
5. Hearing <191>(5723) of thy <4675> love <26> and <2532> faith <4102>, which <3739> thou hast <2192>(5719) toward <4314> the Lord <2962> Jesus <2424>, and <2532> toward <1519> all <3956> saints <40>;
6. That <3704> the communication <2842> of thy <4675> faith <4102> may become <1096>(5638) effectual <1756> by <1722> the acknowledging <1922> of every <3956> good thing <18> which <3588> is in <1722> you <5213> in <1519> Christ <5547> Jesus <2424>.
7. For <1063> we have <2192>(5719) great <4183> joy <5485> and <2532> consolation <3874> in <1909> thy <4675> love <26>, because <3754> the bowels <4698> of the saints <40> are refreshed <373>(5769) by <1223> thee <4675>, brother <80>.
8. Wherefore <1352>, though I might be <2192>(5723) much <4183> bold <3954> in <1722> Christ <5547> to enjoin <2004>(5721) thee <4671> that which is convenient <433>(5723),
9. Yet for <1223> love's <26> sake I <3870> <0> rather <3123> beseech <3870>(5719) thee, being <5607>(5752) such an one <5108> as <5613> Paul <3972> the aged <4246>, and <1161> now <3570> also <2532> a prisoner <1198> of Jesus <2424> Christ <5547>.
10. I beseech <3870>(5719) thee <4571> for <4012> my <1699> son <5043> Onesimus <3682>, whom <3739> I have begotten <1080>(5656) in <1722> my <3450> bonds <1199>:
11. Which <3588> in time past <4218> was <890> <0> to thee <4671> unprofitable <890>, but <1161> now <3570> profitable <2173> to thee <4671> and <2532> to me <1698>:
12. Whom <3739> I have sent again <375>(5656): thou <4771> therefore <1161> receive <4355>(5640) him <846>, that <5123>(5748) is, mine own <1699> bowels <4698>:
13. Whom <3739> I <1473> would <1014>(5711) have retained <2722>(5721) with <4314> me <1683>, that <2443> in thy <4675> stead <5228> he might have ministered <1247>(5725) unto me <3427> in <1722> the bonds <1199> of the gospel <2098>:
14. But <1161> without <5565> thy <4674> mind <1106> would <2309>(5656) I do <4160>(5658) nothing <3762>; that <3363> <0> thy <4675> benefit <18> should <5600> <0> not <3363> be <5600>(5753) as <5613> it were of <2596> necessity <318>, but <235> willingly <1595> <2596>.
15. For <1063> <1223> perhaps <5029> he <5563> <0> therefore <5124> departed <5563>(5681) for <4314> a season <5610>, that <2443> thou shouldest receive <568>(5719) him <846> for ever <166>;
16. Not now <3765> as <5613> a servant <1401>, but <235> above <5228> a servant <1401>, a brother <80> beloved <27>, specially <3122> to me <1698>, but <1161> how much <4214> more <3123> unto thee <4671>, both <2532> in <1722> the flesh <4561>, and <2532> in <1722> the Lord <2962>?
17. If <1487> thou count <2192>(5719) me <1691> therefore <3767> a partner <2844>, receive <4355>(5640) him <846> as <5613> myself <1691>.
18. If <1161> <1487> he hath wronged <91>(5656) thee <4571>, or <2228> oweth <3784>(5719) thee ought <5100>, put <1677> <0> that <5124> on <1677>(5720) mine account <1698>;
19. I <1473> Paul <3972> have written <1125>(5656) it with mine own <1699> hand <5495>, I <1473> will repay <661>(5692) it: albeit <3363> <0> I do <3004> <0> not <3363> say <3004>(5725) to thee <4671> how <3754> thou owest <4359> <0> unto me <3427> even <2532> thine own self <4572> besides <4359>(5719).
20. Yea <3483>, brother <80>, let <3685> <0> me <1473> have joy <3685>(5636) of thee <4675> in <1722> the Lord <2962>: refresh <373>(5657) my <3450> bowels <4698> in <1722> the Lord <2962>.
21. Having confidence <3982>(5756) in thy <4675> obedience <5218> I wrote <1125>(5656) unto thee <4671>, knowing <1492>(5761) that <3754> thou wilt <4160> <0> also <2532> do <4160>(5692) more than <5228> I say <3004>(5719).
22. But <1161> withal <260> prepare <2090>(5720) me <3427> also <2532> a lodging <3578>: for <1063> I trust <1679>(5719) that <3754> through <1223> your <5216> prayers <4335> I shall be given <5483>(5701) unto you <5213>.
23. There salute <782>(5736) thee <4571> Epaphras <1889>, my <3450> fellowprisoner <4869> in <1722> Christ <5547> Jesus <2424>;
24. Marcus <3138>, Aristarchus <708>, Demas <1214>, Lucas <3065>, my <3450> fellowlabourers <4904>.
25. The grace <5485> of our <2257> Lord <2962> Jesus <2424> Christ <5547> be with <3326> your <5216> spirit <4151>. Amen <281>. Written <1125>(5648) from <575> Rome <4516> to <4314> Philemon <5371>, by <1223> Onesimus <3682> a servant <3610>.


![]() Comments made directly on this verse
Comments made directly on this verse ![]() Comments made elsewhere with reference to this verse
Comments made elsewhere with reference to this verse![]() Comments made directly on this verse
Comments made directly on this verse ![]() Comments made elsewhere with reference to this verse
Comments made elsewhere with reference to this verse![]() Comments made directly on this verse
Comments made directly on this verse ![]() Comments made elsewhere with reference to this verse
Comments made elsewhere with reference to this verse loading...
loading...














